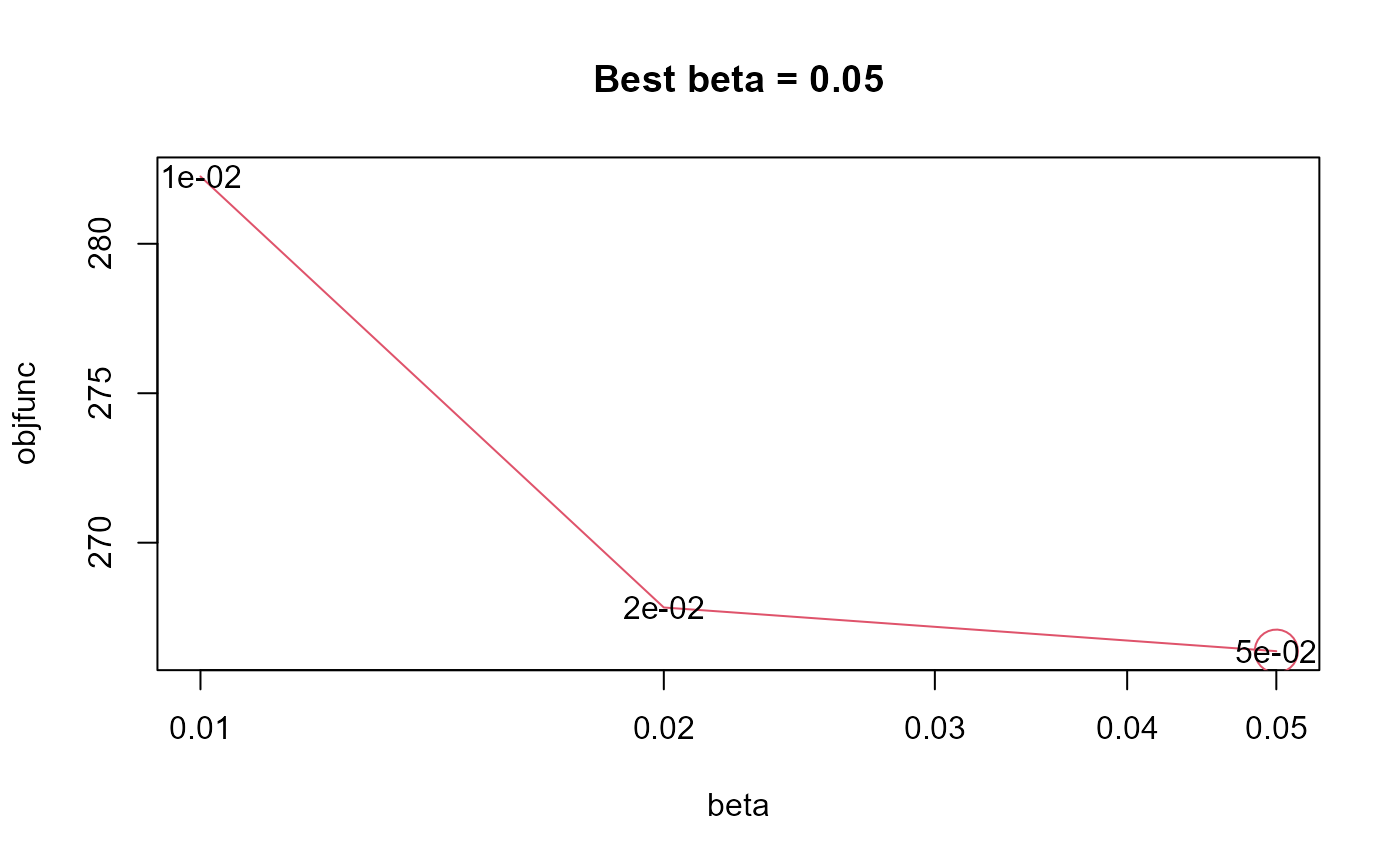
nmfae.kernel.beta.cv selects the optimal beta parameter of the
kernel function by evaluating nmfae.cv for each candidate value.
The kernel matrix \(A = K(U, V; \beta)\) replaces \(Y_2\) in the three-layer
NMF model.
When beta = NULL, candidate values are automatically generated via
nmfkc.kernel.beta.nearest.med.
Usage
nmfae.kernel.beta.cv(
Y1,
rank = 2,
rank.encoder = rank,
U,
V = NULL,
beta = NULL,
plot = TRUE,
...
)Arguments
- Y1
Output matrix \(Y_1\) (P1 x N). Non-negative.
- rank
Integer. Rank of the decoder basis. Default is 2.
- rank.encoder
Integer. Rank of the encoder basis. Default is
rank.- U
Covariate matrix \(U\) (K x M). Rows are features, columns are samples (or knot points for non-symmetric kernels).
- V
Covariate matrix \(V\) (K x N). If
NULL(default),V = Uand a symmetric kernel is used.- beta
Numeric vector of candidate beta values. If
NULL, automatically determined vianmfkc.kernel.beta.nearest.med.- plot
Logical. If
TRUE(default), plots the objective function curve.- ...
Additional arguments. Kernel-specific args (
kernel,degree) are passed tonmfkc.kernel; all others (div,seed,shuffle,epsilon,maxit, etc.) are passed tonmfae.cv. For backward compatibility,QandRare accepted as aliases forrankandrank.encoder.
Value
A list with components:
- beta
The beta value that minimizes the cross-validation objective.
- objfunc
Named numeric vector of objective function values for each candidate beta.