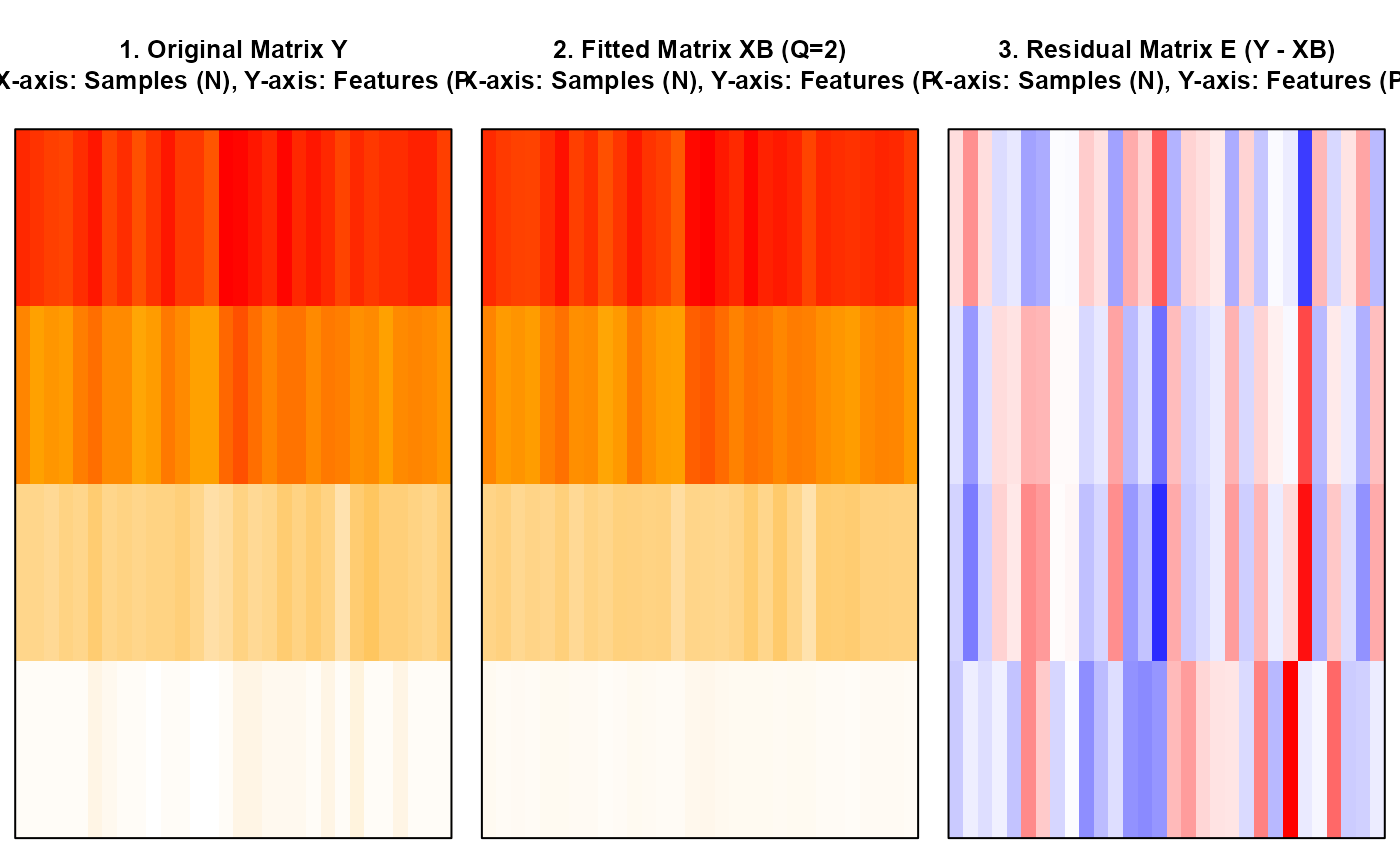
Plot Diagnostics: Original, Fitted, and Residual Matrices as Heatmaps
Source:R/nmfkc.R
nmfkc.residual.plot.RdThis function generates a side-by-side plot of three heatmaps: the original observation matrix Y, the fitted matrix XB (from NMF), and the residual matrix E (Y - XB). This visualization aids in diagnosing whether the chosen rank Q is adequate by assessing if the residual matrix E appears to be random noise.
The axis labels (X-axis: Samples, Y-axis: Features) are integrated into the main title of each plot to maximize the plot area, reflecting the compact layout settings.
Usage
nmfkc.residual.plot(
Y,
result,
fitted.palette = (grDevices::colorRampPalette(c("white", "orange", "red")))(256),
residual.palette = (grDevices::colorRampPalette(c("blue", "white", "red")))(256),
...
)Arguments
- Y
The original observation matrix (P x N).
- result
The result object returned by the nmfkc function.
- fitted.palette
A vector of colors for Y and XB heatmaps. Defaults to white-orange-red. For backward compatibility,
Y_XB_paletteis accepted via....- residual.palette
A vector of colors for the residuals heatmap. Defaults to blue-white-red. For backward compatibility,
E_paletteis accepted via....- ...
Additional graphical parameters passed to the internal image calls.